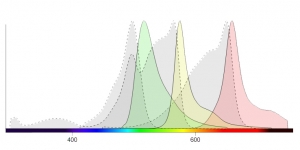
Fluorescence Spectra Viewer Visualize the excitation and emission spectra for popular dyes
Our partner company Biotium is specialized in developing novel fluorescent dyes and products to drive breakthroughs in life science and medical research. Use Biotium's fluorescence spectra viewer to visualize the excitation and emission spectra for their popular CF® Dyes, organelle stain...
The Human Protein Atlas All the human proteins mapped in cells, tissues and organs
The Human Protein Atlas is a Swedish-based program initiated in 2003 with the aim to map all the human proteins in cells, tissues and organs using an integration of various omics technologies, including antibody-based imaging, mass spectrometry-based proteomics, transcriptomics and systems biology. All the data in the knowledge resource is open access to allow scientists both in academia and industry to freely access the data for exploration of the human proteome. The Human Protein Atlas consists of six separate parts, each focusing on a partic...
dianova - Secondary Antibody Finder Select the right secondary antibody in 6 easy steps
dianova is the portal for secondary antibodies and conjugates from BIOZOL. In our antibody search, you can select the properties you need by choosing from 6 properties in drop-down menus, allowing you to quickly and easily find the products best suited to your needs. With more than 7000 secondary antibod...
GenScript Gene Mutagenesis designer Tool to design point DNA mutagenesis to facilitate gene mutation
Gene Mutagenesis Designer is developed to make your design of point DNA mutagenesis straightforward to facilitate gene mutation. To perform DNA mutagenesis from wild type, simply input your starting sequence of wild type gene, and then click on the "from selection" button to select the amino acid(s) of interest. Consequently, the new gene sequence encoding mutated protein will be generated upon a cli...
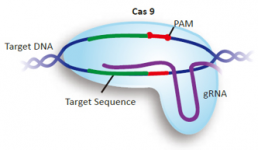
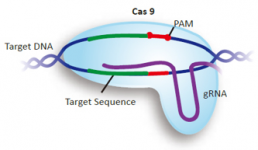
GenScript Genome-wide gRNA databases Genome-wide gRNA databases for CRISPR applications
To facilitate your research using CRISPR technology, GenScript provides genome-wide databases containing pre-validated gRNA sequences design by the Broad Institute of Harvard and MIT. Reference: Sanjana et al., Nat Methods. 2014 Aug;11(8):783-4. doi: 10.1038/nmeth.3047. More than 20,000 gRNA construct...
GenScript Restriction Enzyme Map Analysis Tools This online tool helps you analyze restriction enzyme cutting maps
Restriction digestion also called restriction endonuclease is a process in which DNA is cut at specific sites, dictated by the surrounding DNA sequence. Restriction digestion is accomplished by incubation of the target DNA molecule with restriction enzymes - enzymes that recognize and bind specific DNA sequences and cleave at specific nucleotides either within the recognition sequence or outside of the recognition sequence....
GenScript Restriction Enzyme Tool Search for restriction enzymes by name, recognition sequence or overha...
The Tool allows you to search for restriction enzymes by name, recognition sequence or overhang. Enter your sequence using single letter code nomenclature, and the tool will identify the right enzyme for the job. If the enzyme has isoschizomers (enzymes with the same recognition sequence and cut site) or neoschizomers (enzymes with the same recognition sequence but a different cut site), a list of these enzymes is provided....
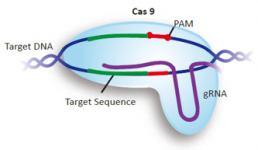
GenScript CRISPR sgRNA Design Tool The design tool will identify single guide RNAs
GenScript is proud to offer free online access to our gRNA sequence design tool, developed by the Broad Institute of Harvard and MIT. Our gRNA design tool will identify single guide RNAs for use with wild-type S. pyogenes Cas9 for any DNA sequence you input. Start your gRNA design project by entering a sequence up to 250bp in length.